AMICI Python example “Boehm”¶
This is an example using the model [boehm_ProteomeRes2014.xml] model to demonstrate and test SBML import and AMICI Python interface.
[1]:
import libsbml
import importlib
import amici
import pypesto
import os
import sys
import numpy as np
import matplotlib.pyplot as plt
# temporarily add the simulate file
sys.path.insert(0, 'boehm_JProteomeRes2014')
from benchmark_import import DataProvider
# sbml file
sbml_file = 'boehm_JProteomeRes2014/boehm_JProteomeRes2014.xml'
# name of the model that will also be the name of the python module
model_name = 'boehm_JProteomeRes2014'
# output directory
model_output_dir = 'tmp/' + model_name
The example model¶
Here we use libsbml to show the reactions and species described by the model (this is independent of AMICI).
[2]:
sbml_reader = libsbml.SBMLReader()
sbml_doc = sbml_reader.readSBML(os.path.abspath(sbml_file))
sbml_model = sbml_doc.getModel()
dir(sbml_doc)
print(os.path.abspath(sbml_file))
print('Species: ', [s.getId() for s in sbml_model.getListOfSpecies()])
print('\nReactions:')
for reaction in sbml_model.getListOfReactions():
reactants = ' + '.join(['%s %s'%(int(r.getStoichiometry()) if r.getStoichiometry() > 1 else '', r.getSpecies()) for r in reaction.getListOfReactants()])
products = ' + '.join(['%s %s'%(int(r.getStoichiometry()) if r.getStoichiometry() > 1 else '', r.getSpecies()) for r in reaction.getListOfProducts()])
reversible = '<' if reaction.getReversible() else ''
print('%3s: %10s %1s->%10s\t\t[%s]' % (reaction.getId(),
reactants,
reversible,
products,
libsbml.formulaToL3String(reaction.getKineticLaw().getMath())))
/home/paul/Documents/pesto/pyPESTO/doc/example/boehm_JProteomeRes2014/boehm_JProteomeRes2014.xml
Species: ['STAT5A', 'STAT5B', 'pApB', 'pApA', 'pBpB', 'nucpApA', 'nucpApB', 'nucpBpB']
Reactions:
v1_v_0: 2 STAT5A -> pApA [cyt * BaF3_Epo * STAT5A^2 * k_phos]
v2_v_1: STAT5A + STAT5B -> pApB [cyt * BaF3_Epo * STAT5A * STAT5B * k_phos]
v3_v_2: 2 STAT5B -> pBpB [cyt * BaF3_Epo * STAT5B^2 * k_phos]
v4_v_3: pApA -> nucpApA [cyt * k_imp_homo * pApA]
v5_v_4: pApB -> nucpApB [cyt * k_imp_hetero * pApB]
v6_v_5: pBpB -> nucpBpB [cyt * k_imp_homo * pBpB]
v7_v_6: nucpApA -> 2 STAT5A [nuc * k_exp_homo * nucpApA]
v8_v_7: nucpApB -> STAT5A + STAT5B [nuc * k_exp_hetero * nucpApB]
v9_v_8: nucpBpB -> 2 STAT5B [nuc * k_exp_homo * nucpBpB]
Importing an SBML model, compiling and generating an AMICI module¶
Before we can use AMICI to simulate our model, the SBML model needs to be translated to C++ code. This is done by amici.SbmlImporter.
[3]:
# Create an SbmlImporter instance for our SBML model
sbml_importer = amici.SbmlImporter(sbml_file)
In this example, we want to specify fixed parameters, observables and a parameter. Unfortunately, the latter two are not part of the SBML standard. However, they can be provided to amici.SbmlImporter.sbml2amici as demonstrated in the following.
Constant parameters¶
Constant parameters, i.e. parameters with respect to which no sensitivities are to be computed (these are often parameters specifying a certain experimental condition) are provided as a list of parameter names.
[4]:
constantParameters = {'ratio', 'specC17'}
Observables¶
We used SBML’s `AssignmentRule <http://sbml.org/Software/libSBML/5.13.0/docs//python-api/classlibsbml_1_1_rule.html>`__ as a non-standard way to specify Model outputs within the SBML file. These rules need to be removed prior to the model import (AMICI does at this time not support these Rules). This can be easily done using amici.assignmentRules2observables().
In this example, we introduced parameters named observable_* as targets of the observable AssignmentRules. Where applicable we have observable_*_sigma parameters for parameters (see below).
[5]:
# Retrieve model output names and formulae from AssignmentRules and remove the respective rules
observables = amici.assignmentRules2observables(
sbml_importer.sbml, # the libsbml model object
filter_function=lambda variable: variable.getId().startswith('observable_') and not variable.getId().endswith('_sigma')
)
print('Observables:', observables)
Observables: {'observable_pSTAT5A_rel': {'name': '', 'formula': '(100 * pApB + 200 * pApA * specC17) / (pApB + STAT5A * specC17 + 2 * pApA * specC17)'}, 'observable_pSTAT5B_rel': {'name': '', 'formula': '-(100 * pApB - 200 * pBpB * (specC17 - 1)) / (STAT5B * (specC17 - 1) - pApB + 2 * pBpB * (specC17 - 1))'}, 'observable_rSTAT5A_rel': {'name': '', 'formula': '(100 * pApB + 100 * STAT5A * specC17 + 200 * pApA * specC17) / (2 * pApB + STAT5A * specC17 + 2 * pApA * specC17 - STAT5B * (specC17 - 1) - 2 * pBpB * (specC17 - 1))'}}
parameters¶
To specify measurement noise as a parameter, we simply provide a dictionary with (preexisting) parameter names as keys and a list of observable names as values to indicate which sigma parameter is to be used for which observable.
[6]:
sigma_vals = ['sd_pSTAT5A_rel', 'sd_pSTAT5B_rel', 'sd_rSTAT5A_rel']
observable_names = observables.keys()
sigmas = dict(zip(list(observable_names), sigma_vals))
print(sigmas)
{'observable_pSTAT5A_rel': 'sd_pSTAT5A_rel', 'observable_pSTAT5B_rel': 'sd_pSTAT5B_rel', 'observable_rSTAT5A_rel': 'sd_rSTAT5A_rel'}
Generating the module¶
Now we can generate the python module for our model. amici.SbmlImporter.sbml2amici will symbolically derive the sensitivity equations, generate C++ code for model simulation, and assemble the python module.
[7]:
sbml_importer.sbml2amici(model_name,
model_output_dir,
verbose=False,
observables=observables,
constantParameters=constantParameters,
sigmas=sigmas
)
Importing the module and loading the model¶
If everything went well, we need to add the previously selected model output directory to our PYTHON_PATH and are then ready to load newly generated model:
[8]:
sys.path.insert(0, os.path.abspath(model_output_dir))
model_module = importlib.import_module(model_name)
And get an instance of our model from which we can retrieve information such as parameter names:
[9]:
model = model_module.getModel()
print("Model parameters:", list(model.getParameterIds()))
print("Model outputs: ", list(model.getObservableIds()))
print("Model states: ", list(model.getStateIds()))
Model parameters: ['Epo_degradation_BaF3', 'k_exp_hetero', 'k_exp_homo', 'k_imp_hetero', 'k_imp_homo', 'k_phos', 'sd_pSTAT5A_rel', 'sd_pSTAT5B_rel', 'sd_rSTAT5A_rel']
Model outputs: ['observable_pSTAT5A_rel', 'observable_pSTAT5B_rel', 'observable_rSTAT5A_rel']
Model states: ['STAT5A', 'STAT5B', 'pApB', 'pApA', 'pBpB', 'nucpApA', 'nucpApB', 'nucpBpB']
Running simulations and analyzing results¶
After importing the model, we can run simulations using amici.runAmiciSimulation. This requires a Model instance and a Solver instance. Optionally you can provide measurements inside an ExpData instance, as shown later in this notebook.
[10]:
h5_file = 'boehm_JProteomeRes2014/data_boehm_JProteomeRes2014.h5'
dp = DataProvider(h5_file)
[11]:
# set timepoints for which we want to simulate the model
timepoints = amici.DoubleVector(dp.get_timepoints())
model.setTimepoints(timepoints)
# set fixed parameters for which we want to simulate the model
model.setFixedParameters(amici.DoubleVector(np.array([0.693, 0.107])))
# set parameters to optimal values found in the benchmark collection
model.setParameterScale(2)
model.setParameters(amici.DoubleVector(np.array([-1.568917588,
-4.999704894,
-2.209698782,
-1.786006548,
4.990114009,
4.197735488,
0.585755271,
0.818982819,
0.498684404
])))
# Create solver instance
solver = model.getSolver()
# Run simulation using model parameters from the benchmark collection and default solver options
rdata = amici.runAmiciSimulation(model, solver)
[12]:
# Create edata
edata = amici.ExpData(rdata, 1.0, 0)
# set observed data
edata.setObservedData(amici.DoubleVector(dp.get_measurements()[0][:, 0]), 0)
edata.setObservedData(amici.DoubleVector(dp.get_measurements()[0][:, 1]), 1)
edata.setObservedData(amici.DoubleVector(dp.get_measurements()[0][:, 2]), 2)
# set standard deviations to optimal values found in the benchmark collection
edata.setObservedDataStdDev(amici.DoubleVector(np.array(16*[10**0.585755271])), 0)
edata.setObservedDataStdDev(amici.DoubleVector(np.array(16*[10**0.818982819])), 1)
edata.setObservedDataStdDev(amici.DoubleVector(np.array(16*[10**0.498684404])), 2)
[13]:
rdata = amici.runAmiciSimulation(model, solver, edata)
print('Chi2 value reported in benchmark collection: 47.9765479')
print('chi2 value using AMICI:')
print(rdata['chi2'])
Chi2 value reported in benchmark collection: 47.9765479
chi2 value using AMICI:
47.97654457818761
Run optimization using pyPESTO¶
[14]:
# create objective function from amici model
# pesto.AmiciObjective is derived from pesto.Objective,
# the general pesto objective function class
model.requireSensitivitiesForAllParameters()
solver.setSensitivityMethod(amici.SensitivityMethod_forward)
solver.setSensitivityOrder(amici.SensitivityOrder_first)
objective = pypesto.AmiciObjective(model, solver, [edata], 1)
[15]:
# create optimizer object which contains all information for doing the optimization
optimizer = pypesto.ScipyOptimizer()
optimizer.solver = 'bfgs'
[16]:
# create problem object containing all information on the problem to be solved
x_names = ['x' + str(j) for j in range(0, 9)]
problem = pypesto.Problem(objective=objective,
lb=-5*np.ones((9)), ub=5*np.ones((9)),
x_names=x_names)
[17]:
# do the optimization
result = pypesto.minimize(problem=problem,
optimizer=optimizer,
n_starts=10) # 200
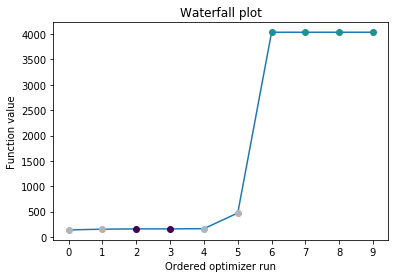
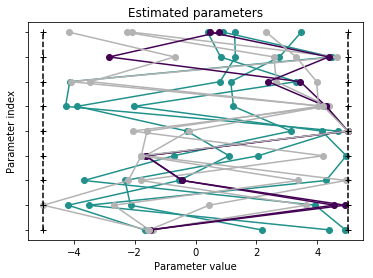
Visualization¶
Create waterfall and parameter plot
[18]:
# waterfall, parameter space,
import pypesto.visualize
pypesto.visualize.waterfall(result)
pypesto.visualize.parameters(result)
[18]:
<matplotlib.axes._subplots.AxesSubplot at 0x7f697f679710>
[ ]: